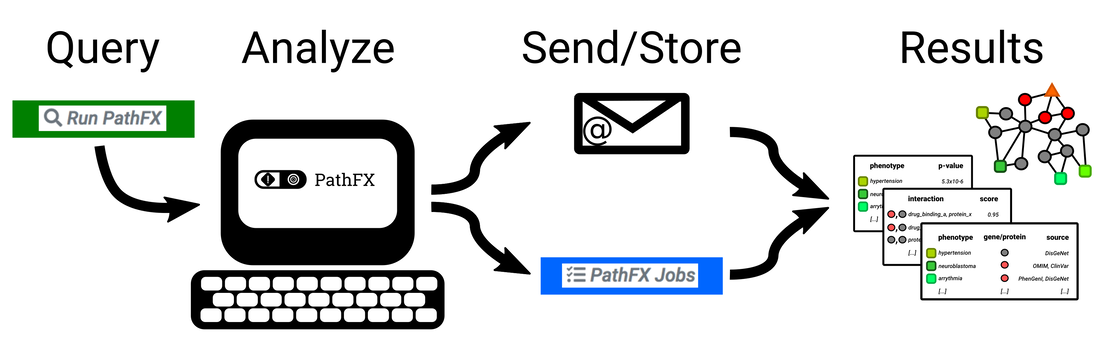
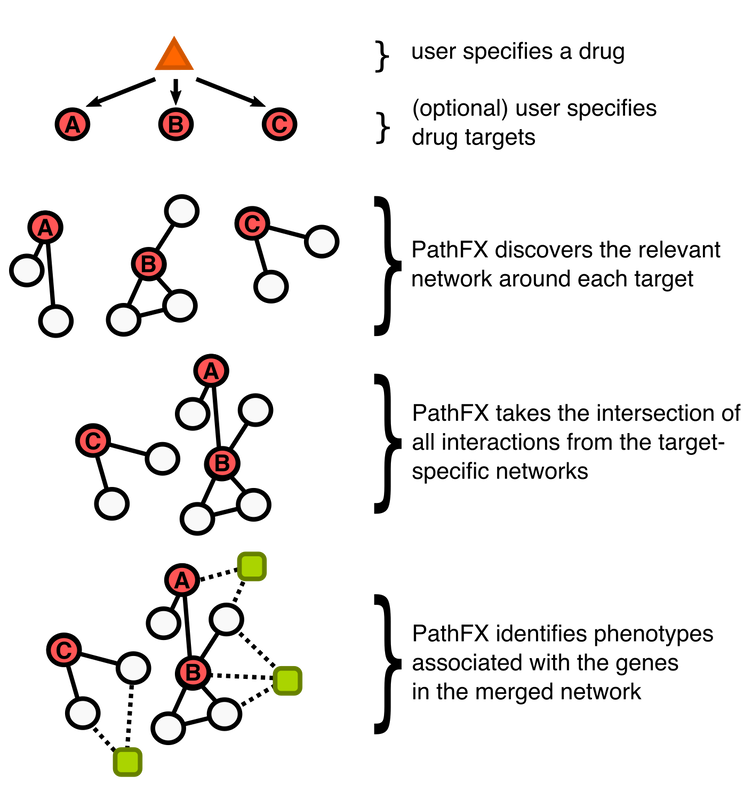
PathFX is designed to look at the pathways-level effects (pathway "FX") of a drug intervention. Under the hood, PathFX is an interaction-network tool that searches for the most relevant protein-protein interactions around a drug's target(s), and then analyzes for which phenotypes the network is enriched relative to the entire interaction network. The algorithm and the web application provide tabular results of associated phenotypes, and a summary figure and table of phenotype clusters based on semantic similarity between phenotypes.
For more information about the PathFX algorithm, please read our paper or visit the code release on GitHub. You can also use the web server application version of PathFX here.
Wilson JL, Racz R, Liu T, Adeniyi O, Sun J, Ramamoorthy A, Pacanowski M, Altman RB. PathFX provides mechanistic insights into drug efficacy and safety for regulatory review and therapeutic development. PLoS computational biology. PLoS Comput Biol. 2018; 14(12): e1006614. PDF Link
Wilson, J.L., M. Wong, A. Chalke, N. Stepanov, D. Petkovic, R.B. Altman. “PathFXweb: a web application for identifying drug safety and efficacy phenotypes.” Bioinformatics. 2019 May 22. pii: btz419. doi: 10.1093/bioinformatics/btz419. [Epub ahead of print]. PDF link Supplement Link
For more information about the PathFX algorithm, please read our paper or visit the code release on GitHub. You can also use the web server application version of PathFX here.
Wilson JL, Racz R, Liu T, Adeniyi O, Sun J, Ramamoorthy A, Pacanowski M, Altman RB. PathFX provides mechanistic insights into drug efficacy and safety for regulatory review and therapeutic development. PLoS computational biology. PLoS Comput Biol. 2018; 14(12): e1006614. PDF Link
Wilson, J.L., M. Wong, A. Chalke, N. Stepanov, D. Petkovic, R.B. Altman. “PathFXweb: a web application for identifying drug safety and efficacy phenotypes.” Bioinformatics. 2019 May 22. pii: btz419. doi: 10.1093/bioinformatics/btz419. [Epub ahead of print]. PDF link Supplement Link
Proudly powered by Weebly